Abstract
Aims/hypothesis
Adipose tissue is a dynamic endocrine organ that regulates whole-body energy homeostasis through the secretion of signalling molecules. Recent reports suggest that secreted microRNAs (miRNAs) may function as biologically active molecules for intercellular communication. This study aims to identify obesity-related circulating miRNA that could be secreted from adipocytes and to explore its possible role in the pathogenesis of metabolic diseases.
Methods
Real-time RT-PCR was used to evaluate the circulating level of miR-130b in mouse models of obesity as well as in humans. Luciferase assay and immunoblotting were used to verify the miRNA target. The effect of miR-130b on mouse peroxisome proliferator-activated receptor γ coactivator-1α was also investigated by electrogene transfer.
Results
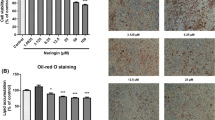
The circulating level of miR-130b was elevated in mouse models of obesity as well as in obese Chinese individuals. More interestingly, the circulating level of miR-130b was positively correlated with BMI. Moreover, circulating miR-130b was a better predictor of the metabolic syndrome than was triacylglycerol level. Mechanistically, adipocytes secreted miR-130b during adipogenesis. TGF-β, which is proportionately increased with obesity, stimulated miR-130b secretion from adipocytes. Furthermore, miR-130b was able to target muscle cells and reduce the expression of its direct target gene, PGC-1α (also known as PPARGC1A), which plays a key role in lipid oxidation in muscle.
Conclusions/interpretation
Circulating miR-130b reflects the degree of obesity and could serve as a potential biomarker for hypertriacylglycerolaemia and metabolic syndrome. Circulating miR-130b could function as a metabolic mediator for adipose–muscle crosstalk and might be involved in the pathogenesis of obesity-associated metabolic diseases.
Similar content being viewed by others
Introduction
Overweight and obesity are defined as abnormal or excessive fat accumulation due to augmented number and size of adipocytes in fat depots. Adipose tissue dysfunction is known to be an important factor in the pathogenesis of type 2 diabetes and other metabolic diseases. In obesity, adipocytes undergo hypertrophy, which leads to adipokine secretion. An increasing body of evidence indicates that dysregulated production of the adipokines participates in the pathogenesis of obesity-associated metabolic disorders [1]. Nowadays, adipose tissue is considered to be a dynamic endocrine organ that regulates nutritional balance and energy homeostasis through crosstalk with other metabolic organs. For example, adipose tissue releases endocrine and metabolic mediators, including TNF-α, IL-6, adiponectin and leptin, to affect insulin signalling and energy metabolism in skeletal muscle.
TGF-β, a key cytokine, regulates adipose tissue metabolism and acts as an adipokine in the pathogenesis of many diseases [2]. Systemic blockade of TGF-β signalling protects mice from obesity, diabetes and hepatic steatosis [3], suggesting that the TGF-β signalling pathway plays an important role in regulating whole-body homeostasis. In humans, circulating TGF-β levels are proportionately increased with obesity in overweight and obese individuals compared with normal-weight individuals [4, 5]. In addition, circulating TGF-β levels, as well as protein levels of TGF-β, in adipocytes are elevated in leptin-deficient (ob/ob) mice and diet-induced obese (DIO) mice [6]. However, the mechanisms underlying the action of TGF-β in the pathogenesis of obesity and obesity-associated metabolic diseases remain largely unknown.
Growing evidence indicates that microRNAs (miRNAs) are key regulators of numerous physiological and pathological processes [7]. During the past several years, studies have demonstrated that miRNAs also exist in various body fluids such as serum, plasma, saliva, urine and milk [8–10]. Altered profiles of circulating miRNAs have been shown to be associated with various diseases, including tissue injury, cancers and diabetes. Consequently, circulating miRNAs have become promising biomarkers for assessing pathophysiological status and monitoring the effectiveness of clinical treatment. More interestingly, it has recently been reported that secreted miRNA could target neighbouring cells or organs and perform a regulatory function in a new location [11, 12]. Although miRNA has been discovered in adipocyte-derived microvesicles [13], and the miRNA expression has been profiled in adipose tissue [14, 15] and the circulation [16, 17] in obese animal models as well as in humans, it remains unclear whether adipocyte-secreted miRNAs are involved in the pathogenesis of obesity-related diseases. Therefore, further investigation is required to determine whether the abnormal circulating miRNA levels could have pathogenic effects.
The objective of this study was to identify a potential miRNA biomarker for obesity and investigate its relationship with obesity-related metabolic diseases and the molecular mechanism involved. Based on our results, we propose that circulating miR-130b could be an early obesity biomarker for clinical investigation and might mediate a metabolic crosstalk between fat and muscle in obesity. Our findings presented here help us to further understand the molecular mechanisms that link obesity and obesity-related metabolic diseases and provide us with new therapeutic targets for defeating obesity-related diseases.
Methods
Reagents, cell culture and differentiation
Adipogenesis of 3T3-L1 pre-adipocytes was induced as described previously with modification [18]. Briefly, on the third day after reaching confluence, 3T3-L1 cells were incubated with 10% FBS, methylisobutylxanthine (115 μg/ml), insulin (1 μg/ml) and dexamethasone (390 ng/ml) for 2 days. Afterwards, cells were maintained in medium containing 10% FBS and insulin (1 μg/ml) for up to 6 days. The culture medium of adipocytes was collected for RNA extraction. Adipocytes were starved of serum for 4 h and then stimulated with 200 nmol/l insulin for 12 h. Myogenesis of C2C12 myoblasts was induced as described previously [19]. Cells were incubated with 5 ng/ml TGF-β (Santa Cruz Biotechnology, Santa Cruz, CA, USA), or as indicated, 50 ng/ml TNF-α (gift from Yingying Le, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, China) and 1 μmol/l rosiglitazone (Cayman Chemical, Ann Arbor, MI, USA) for 12 h. Small interfering RNA (siRNA) oligos and miR-130b mimics were purchased from GenePharma (Shanghai, China).
Mouse and human studies
Mice were maintained and experiments were performed according to the protocol approved by the Animal Care and Use Committees of Institute for Nutritional Sciences (permit No.: 2011-AN-14). ‘Principles of laboratory animal care’ were followed, as well as specific national laws where applicable. Male ob/ob mice, 6–8 weeks old, provided by the Model Animal Research Center of Nanjing University (Nanjing, China), on a C57BL/6 genetic background, were used. To obtain DIO mice, C57BL/6 mice were fed a high-fat diet (HFD; 45% calories from fat) for 4 weeks. Ten- to thirteen-week-old male mice were used in this study. Fat pads were harvested, frozen in liquid nitrogen and stored at −80°C. Blood samples were centrifuged to prepare serum and stored at −80°C.
For the human study, only men were recruited and the study was reviewed and approved by the Ethics Committee of Shanghai Xuhui Central Hospital (Shanghai, China). All participants gave written informed consent. The fasting serum samples were collected at Shanghai Xuhui Central Hospital. The percentage of body fat was estimated by using TANITA-TA-410 body composition analyser (Tanita, Tokyo, Japan). Lipid and glucose measurements were automated by using Siemens Advia 2400 Clinical Chemistry System (Siemens, Erlangen, Germany). Blood pressure was measured by using an Omron blood pressure monitor (Omron, Tokyo, Japan). Human insulin level was measured by using a human INS ELISA Kit (R&D Systems, Minneapolis, MN, USA). The WHO recommends using BMI cut points to define obesity and overweight in adults. In China, BMIs of 24 kg/m2 and 28 kg/m2 are considered to be the best cut points to classify normal weight, overweight and obesity [20].
RNA isolation, quantitative RT-PCR analysis and western blot analysis
The miRNAs were extracted from serum and cell culture medium using the miRcute miRNA isolation kit (Tiangen, Shanghai, China). Total RNA was reverse-transcribed by using the PrimeScript RT reagent Kit (TaKaRa, Shiga, Japan). Small RNA was reverse-transcribed by using stem-loop RT, which was performed according to the protocols [21]. Real-time PCR was performed on an ABI 7900 Real-Time PCR System (Applied Biosystems, Foster City, CA, USA). The expression levels of miRNAs in the cells, tissues, or culture medium were normalised to U6 level. The level of circulating miRNA was normalised to miR-223 level. Western blot analysis was performed as described previously, with modification [18]. Briefly, cultured cells were homogenised in RIPA lysis buffer (50 mmol/l Tris HCl, pH 7.5, 150 mmol/l NaCl, 1.0 mmol/l EDTA, 0.1% wt/vol. SDS, 1% wt/vol. sodium deoxycholate and 1% wt/vol. Triton X-100), which was supplemented with Protease Inhibitor Cocktail and Phosphatase Inhibitor Cocktail (Sigma, St Louis, MO, USA) immediately before use. The protein concentration was determined using a BCA protein assay kit (Thermo Fisher, Waltham, MA, USA). Protein lysates were resolved on 10% SDS-PAGE gels using standard procedures. Anti-PGC-1α (Santa Cruz Biotechnology, Santa Cruz, CA, USA) and anti-GAPDH (Kang Chen, Shanghai, China) were used for western blot analysis.
Plasmid and RNA oligonucleotide
Primers used in the current study are listed in electronic supplementary material (ESM) Table 1. For construction of the luciferase reporter plasmid, the 3′UTR of PGC-1α (also known as PPARGC1A) was amplified from human cDNA by PCR, and inserted into the pRL-TK vector. A mutant derivative (PGC-1α 3′UTR mutation) was obtained by using the QuikChange Lightning Site-Directed Mutagenesis Kit (Stratagene, La Jolla, CA, USA). siRNA information is provided in ESM Table 1.
Transfection and luciferase assay
Transfection was performed using Lipofectamine 2000 (Invitrogen, Carlsbad, CA, USA). Luciferase assays were performed by using the Dual-Luciferase Reporter Assay System (Promega, Madison, WI, USA) as described previously [22]. Luciferase activity was measured on a luminometer (Berthold Technologies, Bad Wildbad, Germany). Electrogene transfer was performed as described previously [23]. Briefly, mice soleus (SOL) muscles were injected with 100 μmol miR-130b mimics. Five 50 V/cm pulses, with a 200 ms interval, were applied. Muscles were collected 6 days after gene delivery.
Prediction of miRNA targets and bioinformatic analysis
miR-130b target genes were predicted by TargetScan (http://www.targetscan.org/). The association between potential miR-130b target genes and human diseases was analysed using the web tool FunDO (http://django.nubic.northwestern.edu/fundo/, accessed July 2012). Only miR-130b target genes associated with metabolic diseases with a p value <0.01 were selected for plotting and listed in a table. The p value was calculated using Fisher’s exact test. The web-based computational tool from DIANA Lab (www.microrna.gr/miRPathv2, accessed July 2012) was used to identify signalling pathways potentially altered by miR-130b targets.
Statistical analysis
Data are presented as means ± SE. For two-group comparisons, Student’s t test was performed; p < 0.05 was considered to represent a significant difference. Receiver operating characteristic (ROC) curves were plotted to evaluate the diagnostic effects of the profiles. The AUC for the ROC curve was calculated for the measurement of discrimination accuracy. GraphPad Prism 5.0 (GraphPad Software, La Jolla, CA, USA) was used for all the statistical analysis.
Results
Circulating levels of miR-130b are significantly elevated in obese mice
We hypothesised that adipose tissue can secret functional miRNAs to crosstalk with other metabolic organs, thereby regulating whole-body metabolism. To test our hypothesis, we carried out experiments to identify miRNAs that are expressed in adipose tissue, dysregulated in obesity and detectable in circulation. We first profiled miRNAs expressed differentially among mouse inguinal white adipose tissue (IWAT), epididymal white adipose tissue (EWAT), brown adipose tissue (BAT), gastrocnemius (GAS) muscle and SOL muscle by microarray, and found that Mir130b expression was relatively higher in white adipose tissue (WAT; ESM Fig. 1). To discover whether Mir130b expression was altered in obesity, we determined its expression in various tissues from ob/ob mice. As shown in Fig. 1a, Mir130b expression was increased dramatically in the EWAT of ob/ob mice compared with that of the wild-type group, which is consistent with an early report [14]. We also observed a slight but significant increase of Mir130b expression in GAS muscle of ob/ob mice compared with wild-type mice.
miR-130b levels are elevated in the WAT and serum of obese mice. (a) Relative expression of Mir130b in the brain (BRA), heart (HEA), kidney (KID), pancreas (PAN), stomach (STO), EWAT, GAS muscle and liver (LIV) of ob/ob mice (black bars) and wild-type mice (white bars). n = 3–6, *p < 0.05 and **p < 0.01 compared with wild-type mice. (b) Relative level of miR-130b in the serum of ob/ob mice as compared with wild-type mice (WT). n = 3–6, ***p < 0.001 compared with WT. (c) Relative expression of Mir130b in the EWAT of DIO mice that were on an HFD and control mice fed a chow diet. n = 4–6, *p < 0.05 compared with control. (d) Relative level of miR-130b in the serum of DIO mice as compared with their controls. n = 3–6, *p < 0.05 compared with control
To determine whether miR-130b also exists in serum and whether the serum levels of miR-130b are altered in obesity, we measured the circulating level of miR-130b in ob/ob mice. Interestingly, we found that the level of miR-130b was significantly elevated in the serum of ob/ob mice (Fig. 1b). A similar elevation of miR-130b level was observed in both EWAT and serum of DIO mice that were on an HFD (Fig. 1c, d). These exciting observations prompted us to investigate the serum level of miR-130b in human samples.
The serum level of miR-130b is positively correlated with obesity in humans
Anthropometric and metabolic characteristics of study participants are summarised in ESM Table 2. As we expected, miR-130b levels were elevated in both overweight and obese participants. Interestingly, a positive correlation between miR-130b level and BMI was evident (Fig. 2a; n = 44, r 2 = 0.6022, p < 0.0001), suggesting a link between the degree of obesity and miR-130b level. Moreover, serum levels of miR-130b could detect individuals who were overweight/obese (BMI ≥ 24 kg/m2) with 70% sensitivity at 95% specificity (Fig. 2b). The ROC curves of miR-130b serum level reflected strong separation between normal and overweight/obese groups, with an AUC of 0.905 (p < 0.0001) (Fig. 2c). Taken together, these results demonstrated that the level of circulating miR-130b is a marker of obesity.
The circulating level of miR-130b is positively correlated with obesity in humans. (a) Circulating level of miR-130b is positively correlated with BMI in humans; n = 44, r 2 = 0.6022, p < 0.0001. (b) Sensitivity and specificity of serum miR-130b level in the diagnosis of overweight/obesity (BMI ≥ 24 kg/m2). The dashed line indicates a 95% specificity threshold. Serum levels of miR-130b could detect overweight/obese individuals with 70% sensitivity. (c) A ROC curve was drawn with the data on serum miR-130b from 44 participants; AUC 0.905, p < 0.0001. (d, e) Circulating miR-130b displays higher sensitivity and specificity for diagnosis of the metabolic syndrome than BF%-BIA. Sensitivity and specificity of serum miR-130b level (d) and BF%-BIA (e) in the diagnosis of metabolic syndrome (MetS). The dashed line indicates a 96% specificity threshold. Serum level of miR-130b could detect individuals with metabolic syndrome with 55% sensitivity (d), while BF%-BIA was less sensitive (30%) (e). The relative level of circulating miR-130b was normalised to miR-223 level (a, b, d)
Circulating miR-130b is a risk factor for metabolic syndrome
To determine whether the circulating level of miR-130b is correlated with other clinical characteristics, correlation analysis was performed. As listed in Table 1, serum levels of miR-130b were also correlated well with body fat percentage estimated by bioelectrical impedance analysis (BF%-BIA), arm circumference, triceps skinfold, calf circumference, waist circumference and triacylglycerol level (TG). These results suggested that the elevated miR-130b levels might associate with poor metabolic profile in humans. In addition, ROC analysis was performed to discover whether miR-130b could be used as a potential biomarker for total cholesterol, TG, hypertension, hyperglycaemia and metabolic syndrome [24]. As shown in Table 2, the ROC analysis of serum level of miR-130b reflected strong separation between normal and metabolic syndrome groups (p = 0.0002). Moreover, the ROC analysis of serum level of miR-130b reflected separation between participants with and without hypertriacylglycerolaemia (p = 0.0043). The separation between participants with and without hypertension also tended to be significant (p = 0.07).
Next, we evaluated the differentiating power of circulating miR-130b levels by comparison with that of BF%-BIA or TG. The ROC curve analysis indicated that BF%-BIA was only useful in discriminating the metabolic syndrome group from the control group with an AUC of 0.727, as shown in Table 2 (p = 0.011). Similarly, the ROC curve analysis indicated that TG was useful in discriminating the metabolic syndrome group from the control group with an AUC of 0.739 as shown in Table 2 (p = 0.0074). These data suggested that miR-130b is a better predictor of individuals with metabolic syndrome than TG, while TG is slightly better than BF%-BIA. Furthermore, serum level of miR-130b could detect individuals with metabolic syndrome with 55% sensitivity at 96% specificity (Fig. 2d), while BF%-BIA was less sensitive (30% and 96%, respectively) for metabolic syndrome diagnosis (Fig. 2e). These results suggested that circulating level of miR-130b displays higher sensitivity and specificity for diagnosis of metabolic syndrome than BF%-BIA.
miR-130b is secreted into culture medium during adipogenesis
Since the levels of miR-130b in serum and WAT were elevated in obese mice (Fig. 1), we speculated that the adipose-derived miR-130b might contribute to the high circulating level of miR-130b. To test this possibility, we first determined whether miR-130b was secreted from adipocytes during adipogenesis. 3T3-L1 pre-adipocytes were differentiated into matured adipocytes by using a standard protocol. As we expected, mRNA levels of Pparg and Adipsin (also known as Cfd), two markers of adipocyte differentiation, as well as Pgc-1α, were significantly increased after differentiation (ESM Fig. 2). Consistent with the findings of a previous report [25], the expression level of Mir130b in adipocytes was slightly decreased during adipogenesis (Fig. 3a). In contrast, we found the level of miR-130b in culture medium was significantly increased (Fig. 3b). Our data indicated that miR-130b is released from differentiating adipocytes during adipogenesis.
The secretion of miR-130b from adipocytes is regulated in physiological and pathological processes. (a, b) miR-130b is released from adipocytes during adipogenesis. The level of miR-130b in adipocytes declines gradually during adipogenesis (a) while the relative level of miR-130b in culture medium increases gradually (b). The relative level of miR-130b was analysed by real-time PCR individually at different time points after induction as indicated (D0, day 0; D2, day 2; D6, day 6). *p < 0.05 and **p < 0.01 compared with D0. (c, d) TGF-β, but not rosiglitazone or TNF-α, is able to increase the secretion of miR-130b from matured adipocytes. The effect of TGF-β, rosiglitazone and TNF-α on the level of miR-130b in the culture medium (c) and on the expression of Mir130b in matured adipocytes (d) was examined by using real-time PCR. *p < 0.05 compared with vehicle (Veh). (e, f) The expression of TGF-β (e) and nSMase2 (f) is increased significantly in the EWAT of ob/ob mice. n = 3, *p < 0.05 and **p < 0.01 compared with wild-type (WT) mice. (g) Knockdown of nSMase2 by siRNA reduces the amount of miR-130b in the culture medium of TGF-β-treated adipocytes. ***p < 0.001 compared with control siRNA
TGF-β stimulates the secretion of miR-130b from matured adipocytes
In addition to investigating the secretion of miR-130b during adipogenesis, we also examined some other signalling factors that might have impact on the release of miR-130b from matured adipocytes. PPARγ ligands and cytokines, such as TNF-α and TGF-β, as well as insulin, are believed to exert complex regulatory actions on adipose tissue under various physiological and pathological conditions. Here we found that treatment with rosiglitazone, TNF-α or insulin did not change the level of miR-130b in the culture medium of matured 3T3-L1 adipocytes (Fig. 3c and ESM Fig. 3a). In contrast, addition of TGF-β dramatically elevated the level of miR-130b in the culture medium (Fig. 3c). Concomitant with TGF-β-induced secretion of miR-130b, the intracellular level of miR-130b was slightly decreased (Fig. 3d). Consistent with previous reports, we also found that the mRNA expression levels of Tgf-β (also known as Tgfb1) were increased in WAT of ob/ob mice compared with wild-type mice (Fig. 3e). These data implicated that high levels of TGF-β in obese individuals might contribute to the increased secretion of miR-130b from the WATs. It is worth noting that the mRNA expression of nSMase2 (also known as Smpd3), encoding neutral sphingomyelinase 2, a regulator of miRNA secretion [26], was also increased in WAT of these obese mice, suggesting the miRNA secretion process is activated in obesity (Fig. 3f). Moreover, TGF-β-induced secretion of miR-130b from adipocytes could be attenuated by knockdown of nSMase2 by siRNA (Fig. 3g). This result indicated that the secretion of miR-130b stimulated by TGF-β in adipocytes is dependent on nSMase2.
Interestingly, TGF-β had no effect on the level of miR-130b in the culture medium of the myotubes or on the intracellular miR-130b level (ESM Fig. 3b, c). Moreover, TGF-β did not increase the level of other miRNAs, such as miR-96 and miR-182, in the culture medium of matured adipocytes (ESM Fig. 3d, e). These data suggested that TGF-β might regulate the secretion of miR-130b in a tissue-specific and miRNA-specific manner.
Potential miR-130b target genes are involved in metabolic pathways and diseases
Since miRNA could have multiple targets and have a differential effect on various signalling pathways, to identify molecular pathways potentially altered by miR-130b, we performed analysis by using a web-based computational tool from DIANA Lab. As shown in ESM Table 3, through its target genes miR-130b might be able to regulate the mTOR and adipocytokine signalling pathways, both of which are important for metabolic regulation. In addition, the correlation between potential miR-130b target genes and diseases was analysed by the FunDO web tool. As shown in Fig. 4 and Table 3, predicted miR-130b target genes were associated with diabetes mellitus, obesity, heart failure, hypertension and atherosclerosis. All these results based on computational analysis implied that miR-130b might regulate metabolism under both physiological and pathological conditions.
Predicted miR-130b target genes are associated with human metabolic diseases. The association between potential miR-130b target genes predicted by TargetScan and human diseases was analysed by the FunDO web tool. Predicted miR-130b target genes associated with metabolic diseases, including diabetes mellitus, obesity, heart failure, hypertension and atherosclerosis, are shown
Circulating miR-130b might affect muscle metabolism by targeting peroxisome proliferator-activated receptor γ coactivator-1α
It has been shown that adipose tissue could function as an endocrine organ and crosstalk with other tissues through various mechanisms. Since the expression of Mir130b was elevated in the skeletal muscle of ob/ob mice (Fig. 1a), we speculated that circulating miR-130b could be taken up by muscle cells and influence the metabolism in its new location. To rule out the possibility that the increased level of miR-130b observed in the skeletal muscle of ob/ob mice was due to the activation of transcription of Mir130b, the level of miR-130b precursor was measured. As shown in Fig. 5a, an increased level of miR-130b precursor was found only in adipose tissues but not in the other tissues examined, including muscle tissues of ob/ob mice, indicating that the high circulating miR-130b level might contribute to the increase of miR-130b level in the muscle of ob/ob mice. In addition, we compared the level of matured miR-130b and miR-130b precursor in three metabolic organs (EWAT, GAS muscle and liver) of wild-type mice. We found that the levels of miR-130b, as well as its precursor, were higher in EWAT than in GAS muscle or liver (Fig. 5b). These results further supported the idea that adipose tissue expression of Mir130b might be the main driver of the changes in circulating miR-130b.
miR-130b regulates muscle metabolism by targeting PGC-1α. (a) The level of miR-130b precursor (pre-miR-130b) is increased significantly only in EWAT but not in brain (BRA), heart (HEA), kidney (KID), pancreas (PAN), stomach (STO), GAS muscle or liver (LIV) of ob/ob mice (black bars) compared with wild-type mice (white bars). n = 3–6, *p < 0.05. (b) The relative level of miR-130b and pre-miR-130b in EWAT, GAS muscle and liver (LIV) of wild-type mice. (c) The effect of culture medium from TGF-β-treated adipocytes on the expression of Mir130b in myotubes. Matured 3T3-L1 adipocytes were treated with different concentrations of TGF-β as indicated; afterwards, the culture medium of adipocytes was collected and added to the culture medium of C2C12 myotubes to mimic a co-culture system. The muscle cells were harvested 12 h after addition of the adipocyte-derived culture medium and miR-130b levels were determined by quantitative RT-PCR. *p < 0.05 compared with no TGF-β treatment. (d) miR-130b regulatory element in the 3′UTR of human PGC-1α was identified by Targetscan. MRE-wt, wild-type MRE; MRE-mut, mutated MRE. (e) The effect of miR-130b mimics on a reporter containing PGC-1α-3′UTR with MRE-wt or MRE-mut was determined in HEK-293T cells. Relative luciferase units (RLU) are shown. White bars, negative control; black bars, miR-130b mimics. Error bars represent the SE of three independent experiments. *p < 0.05 compared with control. (f) Mir130b expression in C2C12 myotubes was determined by quantitative RT-PCR 24 h post-transfection of miR-130b mimics. White bars, negative control; black bars, miR-130b mimics. n = 3, **p < 0.01 compared with control. (g) The effect of miR-130b mimics on PGC-1α protein expression was evaluated by western blot analysis in C2C12 myotubes. Total protein was harvested 24 h or 18 h post-transfection, respectively. NC, negative control. (h) Cpt1b, a PGC-1α target gene, is repressed by miR-130b mimics in C2C12 myotubes. Total RNA was harvested 24 h post-transfection and real-time PCR analysis was performed. White bars, negative control; black bars, miR-130b mimics. Error bars represent the SE of three independent experiments. *p < 0.05 compared with control. (i) The in vivo transfection efficiency was evaluated by quantitative RT-PCR for miR-130b. SOL, soleus muscle. White bars, negative control; black bars, miR-130b mimics. n = 3, ***p < 0.001 compared with control. (j) The effect of miR-130b on PGC-1α protein expression was evaluated by western blot analysis in the soleus muscle of mice. NC, negative control. (k) The effect of miR-130b on the mRNA expression of PGC-1α and Cpt1b was evaluated by real-time PCR analysis in the soleus muscle of mice. White bars, negative control; black bars, miR-130b mimics. n = 3, *p < 0.05 compared with control
To provide direct evidence that miR-130b secreted by adipocytes could be taken up by muscle cells, we treated the matured 3T3-L1 adipocytes with different concentrations of TGF-β; afterwards, the culture medium of adipocytes was collected and added to the culture medium of C2C12 myotubes to mimic a co-culture system. As we expected, the culture medium from the adipocytes treated with TGF-β elevated the miR-130b levels in myotubes in a dose-dependent manner (Fig. 5c), suggesting that miR-130b could be taken up by muscle cells and act as a mediator for adipose–muscle crosstalk.
Among the predicted miR-130b targets, PGC-1α is known to be a crucial regulator for multiple metabolic processes, including lipid oxidation, mitochondrial biogenesis and respiration and fibre-type switching in muscle [27, 28]. To confirm that PGC-1α is a real target of miR-130b, we first cloned 3′UTR of PGC-1α containing the putative miRNA regulatory element (MRE) for miR-130b (Fig. 5d) into a luciferase reporter plasmid to determine whether miR-130b could have inhibitory effect. As shown in Fig. 5e, miR-130b significantly suppressed the luciferase activity suggesting that miR-130b could function through the 3′UTR of PGC-1α to inhibit the reporter gene expression. In addition, we made a luciferase reporter with a mutation in the MRE for miR-130b (Fig. 5d). This mutation abrogated the suppressive effect of miR-130b on the 3′UTR of PGC-1α (Fig. 5e). Moreover, we investigated whether miR-130b could affect the expression of peroxisome proliferator-activated receptor γ coactivator-1α (PGC-1α) in cells. Since PGC-1α expression is abundant in muscles, we examined the effect of miR-130b on PGC-1α expression in myotubes. First, we evaluated the transfection efficiency of miR-130b in C2C12 myotubes. As shown in Fig. 5f, Mir130b expression in myotubes was elevated about 300-fold post-transfection. As we expected, the PGC-1α protein expression, as well as the mRNA expression of its downstream target gene Cpt1b (encoding carnitine palmitoyltransferase 1b [CPT1B]), was decreased by Mir130b overexpression (Fig. 5g, h). These findings suggested that PGC-1α is a direct target gene of miR-130b in muscles.
To determine whether miR-130b could affect PGC-1α and CPT1B in vivo, we delivered miR-130b into the SOL muscle of mice as described in Methods. First, we evaluated the transfection efficiency of miR-130b in SOL muscle. As shown in Fig. 5i, the miR-130b level was elevated by more than 30-fold in vivo. In support of our hypothesis, overexpression of Mir130b in SOL muscle led to a decrease in PGC-1α protein level and mRNA level (Fig. 5j, k). Concomitant with the reduced expression of PGC-1α, the mRNA levels of Cpt1b and those genes involved in mitochondrial electron transport and fatty acid β-oxidation, were decreased (Fig. 5k and ESM Fig. 4), suggesting that the oxidative capacity was impaired in the muscle. Based on these findings, we proposed that the repression of PGC-1α and Cpt1b by miR-130b in the skeletal muscle of obese individuals might contribute to the reduction of oxidative capacity and mitochondrial dysfunction, and lead to metabolic disorder.
Discussion
Since secreted miRNAs can be detected in biological fluids such as serum and reflect the physiological status of the cells and organs they originate from, they could potentially serve as predictive and prognostic biomarkers for human diseases. Also, measurement of circulating miRNA levels is believed to be an alternative or complementary diagnostic strategy to biopsy profiling, especially for asymptomatic patients. In this study, our results revealed a novel obesity biomarker, circulating miR-130b, the level of which was positively correlated with the degree of obesity. In addition, the elevated miR-130b levels in overweight participants before the onset of obesity seemed to be associated with a poor metabolic profile, suggesting that miR-130b level could be used for early detection of metabolic syndrome. Indeed, according to our analysis, the level of circulating miR-130b displayed diagnostic power in detecting hypertriacylglycerolaemia and metabolic syndrome. Moreover, the serum miR-130b level was more closely related to the metabolic syndrome than was BF%-BIA. A recent paper by Ortega et al reported that circulating miR-130b levels are reduced in morbidly obese individuals (BMI ≥ 40 kg/m2) [17]. This discrepancy is probably due to the different BMI ranges and patient populations used in these two studies. It also suggests that in morbidly obese individuals, the crosstalk between tissues might be dysregulated. Actually, Ortega et al also detected a slight increase in miR-130b level in obese (40 kg/m2 > BMI ≥ 30 kg/m2) as compared with non-obese individuals (30 kg/m2 > BMI), although the difference was not statistically significant.
Recently, the TGF-β signalling pathway has been shown to play an important role in regulating whole-body energy homeostasis [3]. Our studies suggested that the miR-130b secreted from adipose tissues might mediate the metabolic regulatory action of TGF-β. In addition, we discovered that TGF-β but not the other factors that we examined could stimulate the secretion of miR-130b from adipocytes. TGF-β promoted the secretion of miR-130b from adipocytes but not from myocytes. TGF-β only induced the secretion of miR-130b of the examined miRNAs from adipocytes in this study. These findings suggested that the secretion of a particular miRNA from certain tissues or organs is tightly controlled under both physiological and pathological conditions to coordinate whole-body energy homeostasis.
WAT dysfunction is believed to be an important factor in the pathogenesis of obesity-related metabolic diseases including type 2 diabetes. Dysregulation of miRNA in adipose tissues and miRNA function in adipocytes has been extensively studied and alteration of circulating miRNA profiles in obese patients has been reported recently. Nonetheless, it is still unclear whether miRNAs could be considered to be a novel class of adipokine and provide a new way for adipose tissues to communicate with other organs. Although the serum pool of miR-130b may arise from other tissues not examined in this study, we believe that we revealed for the first time that miR-130b could function as an ‘adipokine’ and mediate the crosstalk between adipose tissues and skeletal muscle.
Reduced oxidative capacity in skeletal muscle is known to increase the risk of developing metabolic diseases, including type 2 diabetes. Genes involved in oxidative phosphorylation exhibit reduced expression levels in the skeletal muscle of individuals with type 2 diabetes and those in an impaired glucose tolerance stage. Such alteration might present before the onset of impaired glucose tolerance. PGC-1α plays a key role in the control of oxidative capacity and is suppressed in the skeletal muscle in animal models of type 2 diabetes [29]. We demonstrated that circulating miR-130b was able to decrease PGC-1α mRNA levels in recipient myocytes, which might lead to reduced muscle oxidative capacity and accelerate the pathogenesis of metabolic syndrome or type 2 diabetes.
Taken together, we identified miR-130b as a potential biomarker for overweight, hypertriacylglycerolaemia and metabolic syndrome, and discovered a novel mechanism linking obesity and obesity-related metabolic diseases, which involves circulating miRNA-based adipose–muscle crosstalk (Fig. 6). This work also broadened our awareness of the crosstalk between adipose tissue and skeletal muscle through secreted miRNA.
Abbreviations
- BF%-BIA:
-
Body fat percentage estimated by bioelectrical impedance analysis
- CPT1B:
-
Carnitine palmitoyltransferase 1b
- DIO:
-
Diet-induced obese
- EWAT:
-
Epididymal WAT
- GAS:
-
Gastrocnemius
- HFD:
-
High-fat diet
- miRNA:
-
microRNA
- MRE:
-
miRNA regulatory element
- nSMase:
-
Neutral sphingomyelinase 2
- PGC-1α:
-
Peroxisome proliferator-activated receptor γ coactivator-1α
- ROC:
-
Receiver operating characteristic
- siRNA:
-
Small interfering RNA
- SOL:
-
Soleus
- WAT:
-
White adipose tissue
References
Ahima RS, Lazar MA (2008) Adipokines and the peripheral and neural control of energy balance. Mol Endocrinol 22:1023–1031
Zamani N, Brown CW (2011) Emerging roles for the transforming growth factor-β superfamily in regulating adiposity and energy expenditure. Endocr Rev 32:387–403
Yadav H, Quijano C, Kamaraju AK et al (2011) Protection from obesity and diabetes by blockade of TGF-beta/Smad3 signaling. Cell Metab 14:67–79
Fain JN, Tichansky DS, Madan AK (2005) Transforming growth factor beta1 release by human adipose tissue is enhanced in obesity. Metabolism 54:1546–1551
Alessi MC, Bastelica D, Morange P et al (2000) Plasminogen activator inhibitor 1, transforming growth factor-beta1, and BMI are closely associated in human adipose tissue during morbid obesity. Diabetes 49:1374–1380
Samad F, Yamamoto K, Pandey M, Loskutoff DJ (1997) Elevated expression of transforming growth factor-beta in adipose tissue from obese mice. Mol Med 3:37–48
Filipowicz W, Bhattacharyya SN, Sonenberg N (2008) Mechanisms of post-transcriptional regulation by microRNAs: are the answers in sight? Nat Rev Genet 9:102–114
Chen X, Gao C, Li H et al (2010) Identification and characterization of microRNAs in raw milk during different periods of lactation, commercial fluid, and powdered milk products. Cell Res 20:1128–1137
Michael A, Bajracharya SD, Yuen PS et al (2010) Exosomes from human saliva as a source of microRNA biomarkers. Oral Dis 16:34–38
Zen K, Zhang C-Y (2012) Circulating microRNAs: a novel class of biomarkers to diagnose and monitor human cancers. Med Res Rev 32:326–348
Chen X, Liang H, Zhang J, Zen K, Zhang CY (2012) Horizontal transfer of microRNAs: molecular mechanisms and clinical applications. Protein Cell 3:28–37
Mittelbrunn M, Gutierrez-Vazquez C, Villarroya-Beltri C et al (2011) Unidirectional transfer of microRNA-loaded exosomes from T cells to antigen-presenting cells. Nat Commun 2:282
Ogawa R, Tanaka C, Sato M et al (2010) Adipocyte-derived microvesicles contain RNA that is transported into macrophages and might be secreted into blood circulation. Biochem Biophys Res Commun 398:723–729
Bengestrate L, Virtue S, Campbell M et al (2011) Genome-wide profiling of microRNAs in adipose mesenchymal stem cell differentiation and mouse models of obesity. PLoS One 6:e21305
Keller P, Gburcik V, Petrovic N et al (2011) Gene-chip studies of adipogenesis-regulated microRNAs in mouse primary adipocytes and human obesity. BMC Endocr Disord 11:7
Heneghan HM, Miller N, McAnena OJ, O'Brien T, Kerin MJ (2011) Differential miRNA expression in omental adipose tissue and in the circulation of obese patients identifies novel metabolic biomarkers. J Clin Endocrinol Metab 96:E846–E850
Ortega FJ, Mercader JM, Catalan V et al (2013) Targeting the circulating microRNA signature of obesity. Clin Chem 59:781–792
Ying H, Araki O, Furuya F, Kato Y, Cheng SY (2007) Impaired adipogenesis caused by a mutated thyroid hormone alpha1 receptor. Mol Cell Biol 27:2359–2371
Zhang D, Li X, Chen C et al (2012) Attenuation of p38-mediated miR-1/133 expression facilitates myoblast proliferation during the early stage of muscle regeneration. PLoS One 7:e41478
Wang Y, Mi J, Shan XY, Wang QJ, Ge KY (2007) Is China facing an obesity epidemic and the consequences? The trends in obesity and chronic disease in China. Int J Obes (Lond) 31:177–188
Chen C, Ridzon DA, Broomer AJ et al (2005) Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res 33:e179
Song Y, Shan S, Zhang Y et al (2012) Ligand-dependent corepressor acts as a novel corepressor of thyroid hormone receptor and represses hepatic lipogenesis in mice. J Hepatol 56:248–254
Sandri M, Sandri C, Gilbert A et al (2004) Foxo transcription factors induce the atrophy-related ubiquitin ligase atrogin-1 and cause skeletal muscle atrophy. Cell 117:399–412
Sun L, Franco OH, Hu FB et al (2008) Ferritin concentrations, metabolic syndrome, and type 2 diabetes in middle-aged and elderly Chinese. J Clin Endocrinol Metab 93:4690–4696
Lee EK, Lee MJ, Abdelmohsen K et al (2011) miR-130 suppresses adipogenesis by inhibiting peroxisome proliferator-activated receptor gamma expression. Mol Cell Biol 31:626–638
Kosaka N, Iguchi H, Yoshioka Y, Takeshita F, Matsuki Y, Ochiya T (2010) Secretory mechanisms and intercellular transfer of microRNAs in living cells. J Biol Chem 285:17442–17452
Nikolic N, Rhedin M, Rustan AC, Storlien L, Thoresen GH, Stromstedt M (2012) Overexpression of PGC-1alpha increases fatty acid oxidative capacity of human skeletal muscle cells. Biochem Res Int 2012:714074
Schuler M, Ali F, Chambon C et al (2006) PGC1alpha expression is controlled in skeletal muscles by PPARbeta, whose ablation results in fiber-type switching, obesity, and type 2 diabetes. Cell Metab 4:407–414
Sparks LM, Xie H, Koza RA et al (2005) A high-fat diet coordinately downregulates genes required for mitochondrial oxidative phosphorylation in skeletal muscle. Diabetes 54:1926–1933
Acknowledgements
We thank Ping Wang (Maastricht University, Maastricht, the Netherlands) for critical review of the manuscript.
Funding
This work was supported by grants from Xuhui Central Hospital (Shanghai, China), the Ministry of Science and Technology of China (2009CB919000, 2010CB912500, 2012BAK01B00, 2010CB945100, 2011CB944104), the National Natural Science Foundation (30970587, 31070679, 81172009) and the Fundamental Research Funds for the Central Universities (JY).
Duality of interest
The authors declare that there is no duality of interest associated with this manuscript.
Contribution statement
This study was designed by YW, YL and HY. Cell culture and animal studies were carried out by YL, YW, DZ and HZ. YW, XW, QW, YH, JW, LZ and HX performed the clinical part of the work. YL, YW and HY analysed the data and wrote the paper. JY, XL and HY contributed to discussion and supervised the project. All authors participated in data interpretation and revising the paper and approved the final version of the manuscript.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Yu-cheng Wang and Yuying Li contributed equally to this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM Fig. 1
Tissue expression pattern of miR-130b in mouse. Microarray analysis of Mir130b expression in mouse inguinal white adipose tissue (IWAT), epididymal white adipose tissue (EWAT), brown adipose tissue (BAT), gastrocnemius (GAS) muscle, and soleus (SOL) muscle. Pooled samples from 5 mice were used for microarray analysis. (PDF 21 kb)
ESM Fig. 2
Differentiation of 3T3-L1 preadipocytes into mature adipocytes. (a-c) The mRNA expression of adipocyte-differentiation markers, PPARγ (a), Adipsin (b), and PGC-1α (c) was measured by qRT-PCR analysis at different time points as indicated during adipogenesis of 3T3-L1 preadipocytes. Error bars represent the SE of three independent experiments. *p < 0.05, **p < 0.01, ***p < 0.001. (PDF 59 kb)
ESM Fig. 3
Effect of insulin or TGF-β on miRNA expression or secretion in myotubes or adipocytes. (a) Matured 3T3-L1 adipocytes were stimulated by insulin for 12 h. miR-130b level in culture medium was determined by qRT-PCR. (b-c) C2C12 myotubes were stimulated with TGF-β for 12 h. miR-130b level in culture medium and intracellular Mir130b expression level were determined by qRT-PCR. Error bars represent the SE of three independent experiments. (d-e) Matured 3T3-L1 adipocytes were incubated with TGF-β, rosiglitazone (Rosi), or TNF-α for 12 h. The levels of miR-96 and miR-182 in the culture medium were measured by qRT-PCR. Error bars represent the SE of three independent experiments. *p < 0.05. (PDF 89 kb)
ESM Fig. 4
The effect of miR-130b on the mRNA expression of genes involved in mitochondrial electron transport and fatty acid β-oxidation. The mRNA expression of cytochrome c (Cyt c), mitochondrial transcription factor A (mtTFA), estrogen-related receptor α (ERRα), cytochrome c oxidase subunit IV (COX4) and superoxide dismutase-2 (SOD2) were evaluated by Real time PCR analysis in the soleus muscle of mice. n = 3, **p < 0.01, ***p < 0.001. White bars: negative control; Black bars: miR-130b mimics. (PDF 25 kb)
ESM Table 1
(PDF 48 kb)
ESM Table 2
(PDF 8 kb)
ESM Table 3
(PDF 7 kb)
Rights and permissions
About this article
Cite this article
Wang, Yc., Li, Y., Wang, Xy. et al. Circulating miR-130b mediates metabolic crosstalk between fat and muscle in overweight/obesity. Diabetologia 56, 2275–2285 (2013). https://doi.org/10.1007/s00125-013-2996-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-013-2996-8