Abstract
Purpose
To evaluate an alternative in vitro system which can provide more quantitatively accurate drug drug interaction (DDI) prediction for 10 protein kinase inhibitors for which DDI risk was over-predicted by inhibition data generated in human liver microsomes (HLM).
Methods
Three cryopreserved human hepatocyte (hHEP) systems: 1) plated hHEPs; 2) hHEPs suspended in Dulbecco’s Modified Eagle Medium (DMEM) and 3) hHEPs suspended in human plasma (plasma hHEPs) were developed to detect CYP3A time dependent inhibition, and the static mechanistic model was used to predict clinical outcomes.
Results
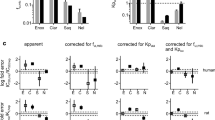
A general trend was observed in the CYP3A inactivation potency (k inact /K I, app ) as HLM > plated > DMEM ≥ plasma hHEPs. Using the static mechanistic model, DDIs predicted using parameters estimated from plated, DMEM and plasma hHEPs had 84, 74 and 95% accuracy (out of 19 clinical interaction studies) within 2-fold of the reported interaction, respectively. They demonstrated significant improvement compared to the DDIs predicted using parameters estimated from HLMs where 58% accuracy was obtained.
Conclusions
Based on 19 DDIs, plasma hHEPs demonstrate a more reliable clinical DDI prediction for 10 protein kinase inhibitors and prototypical CYP3A time dependent inhibitors.





Similar content being viewed by others
Abbreviations
- [I] g :
-
The intestinal inhibitor concentration
- [I] h :
-
The hepatic inhibitor concentrations
- DDI:
-
Drug drug interactions
- DMEM:
-
Dulbecco’s Modified Eagle Medium
- fu,mic :
-
Unbound fraction in human liver microsomes
- fu,hep :
-
The non specific binding in the hepatocyte incubation
- f u,p :
-
Unbound fraction in human plasma
- GMFE:
-
The geometric mean-fold error
- hHEPs:
-
Human hepatocytes
- HLM:
-
Human liver microsomes
- IS:
-
Internal standard
- k deg :
-
Degradation rate constant
- K I :
-
Inactivation constant
- K i :
-
Inhibition constant
- K I,app :
-
Apparent inactivation constant
- K i,app :
-
Apparent inhibition constant
- k inact :
-
Maximum inactivation rate constant
- RMSE:
-
The root-mean-square error
- TDI:
-
Time dependent inhibition
References
Grimm SW, Einolf HJ, Hall SD, He K, Lim HK, Ling KH, et al. The conduct of in vitro studies to address time-dependent inhibition of drug-metabolizing enzymes: a perspective of the pharmaceutical research and manufacturers of America. Drug Metab Dispos Biol Fate Chem. 2009;37(7):1355–70.
Chen Y, Liu L, Monshouwer M, Fretland AJ. Determination of time-dependent inactivation of CYP3A4 in cryopreserved human hepatocytes and assessment of human drug-drug interactions. Drug Metab Dispos Biol Fate Chem. 2011;39(11):2085–92.
Xu L, Chen Y, Pan Y, Skiles GL, Shou M. Prediction of human drug-drug interactions from time-dependent inactivation of CYP3A4 in primary hepatocytes using a population-based simulator. Drug Metab Dispos Biol Fate Chem. 2009;37(12):2330–9.
Kenny JR, Mukadam S, Zhang C, Tay S, Collins C, Galetin A, et al. Drug-drug interaction potential of marketed oncology drugs: in vitro assessment of time-dependent cytochrome P450 inhibition, reactive metabolite formation and drug-drug interaction prediction. Pharm Res. 2012;29(7):1960–76.
Mao J, Johnson TR, Shen Z, Yamazaki S. Prediction of crizotinib-midazolam interaction using the Simcyp population-based simulator: comparison of CYP3A time-dependent inhibition between human liver microsomes versus hepatocytes. Drug Metab Dispos Biol Fate Chem. 2013;41(2):343–52.
Calvert H, Twelves C, Ranson M, Plummer R, Fettner S, Pantze M, et al. Effect of erlotinib on CYP3A activity, evaluated in vitro and by dual probes in patients with cancer. Anti-Cancer Drugs. 2014;25(7):832–40.
Filppula AM, Neuvonen PJ, Backman JT. In vitro assessment of time-dependent inhibitory effects on CYP2C8 and CYP3A activity by fourteen protein kinase inhibitors. Drug Metab Dispos Biol Fate Chem. 2014;42(7):1202–9.
Mao J, Mohutsky MA, Harrelson JP, Wrighton SA, Hall SD. Prediction of CYP3A-mediated drug-drug interactions using human hepatocytes suspended in human plasma. Drug Metab Dispos Biol Fate Chem. 2011;39(4):591–602.
Mao J, Mohutsky MA, Harrelson JP, Wrighton SA, Hall SD. Predictions of cytochrome P450-mediated drug-drug interactions using cryopreserved human hepatocytes: comparison of plasma and protein-free media incubation conditions. Drug Metab Dispos Biol Fate Chem. 2012;40(4):706–16.
Albaugh DR, Fullenwider CL, Fisher MB, Hutzler JM. Time-dependent inhibition and estimation of CYP3A clinical pharmacokinetic drug-drug interactions using plated human cell systems. Drug Metab Dispos Biol Fate Chem. 2012;40(7):1336–44.
Fahmi OA, Maurer TS, Kish M, Cardenas E, Boldt S, Nettleton D. A combined model for predicting CYP3A4 clinical net drug-drug interaction based on CYP3A4 inhibition, inactivation, and induction determined in vitro. Drug Metab Dispos Biol Fate Chem. 2008;36(8):1698–708.
Gertz M, Harrison A, Houston JB, Galetin A. Prediction of human intestinal first-pass metabolism of 25 CYP3A substrates from in vitro clearance and permeability data. Drug Metab Dispos Biol Fate Chem. 2010;38(7):1147–58.
Yang J, Liao M, Shou M, Jamei M, Yeo KR, Tucker GT, et al. Cytochrome p450 turnover: regulation of synthesis and degradation, methods for determining rates, and implications for the prediction of drug interactions. Curr Drug Metab. 2008;9(5):384–94.
Ernest 2nd CS, Hall SD, Jones DR. Mechanism-based inactivation of CYP3A by HIV protease inhibitors. J Pharmacol Exp Ther. 2005;312(2):583–91.
Obach RS, Walsky RL, Venkatakrishnan K, Gaman EA, Houston JB, Tremaine LM. The utility of in vitro cytochrome P450 inhibition data in the prediction of drug-drug interactions. J Pharmacol Exp Ther. 2006;316(1):336–48.
Han B, Dickinson GL, Turner PK, Hall SD. Prediction of CYP3A mediated drug-drug interactions: estimation of gut wall and hepatic contributions. ASCPT 2013 poster; 2013.
Galetin A, Burt H, Gibbons L, Houston JB. Prediction of time-dependent CYP3A4 drug-drug interactions: impact of enzyme degradation, parallel elimination pathways, and intestinal inhibition. Drug Metab Dispos Biol Fate Chem. 2006;34(1):166–75.
Houston JB, Galetin A. Methods for predicting in vivo pharmacokinetics using data from in vitro assays. Curr Drug Metab. 2008;9(9):940–51.
Rostami-Hodjegan A, Tucker G. ‘In silico’ simulations to assess the ‘in vivo’ consequences of ‘in vitro’ metabolic drug–drug interactions. Drug Discov Today. 2004;1(4):441–8.
Duckett DR, Cameron MD. Metabolism considerations for kinase inhibitors in cancer treatment. Expert Opin Drug Metab Toxicol. 2010;6(10):1175–93.
Kilford PJ, Gertz M, Houston JB, Galetin A. Hepatocellular binding of drugs: correction for unbound fraction in hepatocyte incubations using microsomal binding or drug lipophilicity data. Drug Metab Dispos Biol Fate Chem. 2008;36(7):1194–7.
NDA2011. http://www.accessdata.fda.gov/drugsatfda_docs/nda/2011/202570Orig1s000ClinPharmR.pdf.
NDA2006. http://www.accessdata.fda.gov/drugsatfda_docs/nda/2006/021986s000_Sprycel__ClinPharmR.pdf.
Okudaira T, Kotegawa T, Imai H, Tsutsumi K, Nakano S, Ohashi K. Effect of the treatment period with erythromycin on cytochrome P450 3A activity in humans. J Clin Pharmacol. 2007;47(7):871–6.
Zimmermann T, Yeates RA, Laufen H, Scharpf F, Leitold M, Wildfeuer A. Influence of the antibiotics erythromycin and azithromycin on the pharmacokinetics and pharmacodynamics of midazolam. Arzneimittelforschung. 1996;46(2):213–7.
Olkkola KT, Aranko K, Luurila H, Hiller A, Saarnivaara L, Himberg JJ, et al. A potentially hazardous interaction between erythromycin and midazolam. Clin Pharmacol Ther. 1993;53(3):298–305.
NDA2009. http://www.accessdata.fda.gov/drugsatfda_docs/nda/2009/022334s000_ClinPharmR.pdf.
O’Brien SG, Meinhardt P, Bond E, Beck J, Peng B, Dutreix C, et al. Effects of imatinib mesylate (STI571, Glivec) on the pharmacokinetics of simvastatin, a cytochrome p450 3A4 substrate, in patients with chronic myeloid leukaemia. Br J Cancer. 2003;89(10):1855–9.
Milton DT, Riely GJ, Azzoli CG, Gomez JE, Heelan RT, Kris MG, et al. Phase 1 trial of everolimus and gefitinib in patients with advanced nonsmall-cell lung cancer. Cancer. 2007;110(3):599–605.
GSKClinicalStudyRegister. RM2006/00931/00, https://gsk.sylogent.com/files/20663.pdf.
NDA2007. http://www.accessdata.fda.gov/drugsatfda_docs/nda/2007/022068s000_ClinPharmR.pdf.
Goh BC, Reddy NJ, Dandamudi UB, Laubscher KH, Peckham T, Hodge JP, et al. An evaluation of the drug interaction potential of pazopanib, an oral vascular endothelial growth factor receptor tyrosine kinase inhibitor, using a modified Cooperstown 5+1 cocktail in patients with advanced solid tumors. Clin Pharmacol Ther. 2010;88(5):652–9.
Flaherty KT, Lathia C, Frye RF, Schuchter L, Redlinger M, Rosen M, et al. Interaction of sorafenib and cytochrome P450 isoenzymes in patients with advanced melanoma: a phase I/II pharmacokinetic interaction study. Cancer Chemother Pharmacol. 2011;68(5):1111–8.
Phimmasone S, Kharasch ED. A pilot evaluation of alfentanil-induced miosis as a noninvasive probe for hepatic cytochrome P450 3A4 (CYP3A4) activity in humans. Clin Pharmacol Ther. 2001;70(6):505–17.
Kharasch ED, Walker A, Hoffer C, Sheffels P. Intravenous and oral alfentanil as in vivo probes for hepatic and first-pass cytochrome P450 3A activity: noninvasive assessment by use of pupillary miosis. Clin Pharmacol Ther. 2004;76(5):452–66.
Wang YH, Jones DR, Hall SD. Prediction of cytochrome P450 3A inhibition by verapamil enantiomers and their metabolites. Drug Metab Dispos Biol Fate Chem. 2004;32(2):259–66.
Stresser DM, Mao J, Kenny JR, Jones BC, Grime K. Exploring concepts of in vitro time-dependent CYP inhibition assays. Expert Opin Drug Metab Toxicol. 2014;10(2):157–74.
McGinnity DF, Berry AJ, Kenny JR, Grime K, Riley RJ. Evaluation of time-dependent cytochrome P450 inhibition using cultured human hepatocytes. Drug Metab Dispos Biol Fate Chem. 2006;34(8):1291–300.
Dong PP, Fang ZZ, Zhang YY, Ge GB, Mao YX, Zhu LL, et al. Substrate-dependent modulation of the catalytic activity of CYP3A by erlotinib. Acta Pharmacol Sin. 2011;32(3):399–407.
Foti RS, Rock DA, Wienkers LC, Wahlstrom JL. Selection of alternative CYP3A4 probe substrates for clinical drug interaction studies using in vitro data and in vivo simulation. Drug Metab Dispos Biol Fate Chem. 2010;38(6):981–7.
VandenBrink BM, Foti RS, Rock DA, Wienkers LC, Wahlstrom JL. Prediction of CYP2D6 drug interactions from in vitro data: evidence for substrate-dependent inhibition. Drug Metab Dispos Biol Fate Chem. 2012;40(1):47–53.
Hu S, Franke RM, Filipski KK, Hu C, Orwick SJ, de Bruijn EA, et al. Interaction of imatinib with human organic ion carriers. Clin Cancer Res Off J Am Assoc Cancer Res. 2008;14(10):3141–8.
Wang L, Giannoudis A, Lane S, Williamson P, Pirmohamed M, Clark RE. Expression of the uptake drug transporter hOCT1 is an important clinical determinant of the response to imatinib in chronic myeloid leukemia. Clin Pharmacol Ther. 2008;83(2):258–64.
Tang SC, de Vries N, Sparidans RW, Wagenaar E, Beijnen JH, Schinkel AH. Impact of P-glycoprotein (ABCB1) and breast cancer resistance protein (ABCG2) gene dosage on plasma pharmacokinetics and brain accumulation of dasatinib, sorafenib, and sunitinib. J Pharmacol Exp Ther. 2013;346(3):486–94.
Swift B, Pfeifer ND, Brouwer KL. Sandwich-cultured hepatocytes: an in vitro model to evaluate hepatobiliary transporter-based drug interactions and hepatotoxicity. Drug Metab Rev. 2010;42(3):446–71.
Di Gion P, Kanefendt F, Lindauer A, Scheffler M, Doroshyenko O, Fuhr U, et al. Clinical pharmacokinetics of tyrosine kinase inhibitors: focus on pyrimidines, pyridines and pyrroles. Clin Pharmacokinet. 2011;50(9):551–603.
Eechoute K, Sparreboom A, Burger H, Franke RM, Schiavon G, Verweij J, et al. Drug transporters and imatinib treatment: implications for clinical practice. Clin Cancer Res Off J Am Assoc Cancer Res. 2011;17(3):406–15.
Zhao P, Kunze KL, Lee CA. Evaluation of time-dependent inactivation of CYP3A in cryopreserved human hepatocytes. Drug Metab Dispos Biol Fate Chem. 2005;33(6):853–61.
Poulin P, Burczynski FJ, Haddad S. The role of extracellular binding proteins in the cellular uptake of drugs: impact on quantitative in vitro-to-in vivo extrapolations of toxicity and efficacy in physiologically based pharmacokinetic-pharmacodynamic research. J Pharm Sci. 2015. doi:10.1002/jps.24571.
ACKNOWLEDGMENTS AND DISCLOSURES
This study was funded by Genentech (A member of the Roche group). All authors were employees of Genentech when this work was carried out. They have no other conflicts of interest to declare. We would like to thank Dr. Steve Wrighton for his thorough review of this manuscript, and Dr. Jason Halladay for the helpful suggestions.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 106 kb)
Rights and permissions
About this article
Cite this article
Mao, J., Tay, S., Khojasteh, C.S. et al. Evaluation of Time Dependent Inhibition Assays for Marketed Oncology Drugs: Comparison of Human Hepatocytes and Liver Microsomes in the Presence and Absence of Human Plasma. Pharm Res 33, 1204–1219 (2016). https://doi.org/10.1007/s11095-016-1865-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11095-016-1865-9